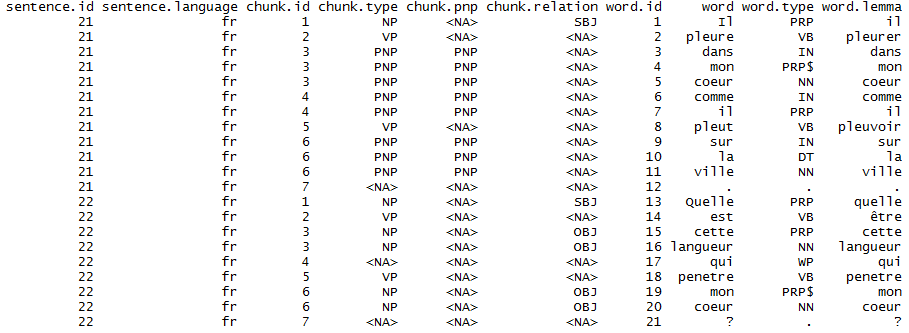
Part of the R course offering of BNOSAC which you can find at http://bnosac.be/images/bnosac/bnosac_courses_r.pdf, we offer several 2-day hands-on courses covering the use of text mining tools for the purpose of data analysis. It covers basic text handling, natural language engineering and statistical modelling on top of textual data.
Interested in upgrading your skills on text mining with R? Registering can be done for the following days.
2016: October 24-25: subscribe at https://lstat.kuleuven.be/training/coursedescriptions/text-mining-with-r
2016: November 14-15: subscribe at http://di-academy.com/event/text-mining-with-r/
2017: March 23-24: subscribe at https://lstat.kuleuven.be/training/coursedescriptions/text-mining-with-r
The following elements are covered in this course.
- Import of (structured) text data with focus on text encodings. Detection of language
- Cleaning of text data, regular expressions
- String distances
- Graphical displays of text data
- Natural language processing: stemming, parts-of-speech (POS) tagging, tokenization, lemmatisation, entity recognition
- Sentiment analysis
- Statistical topic detection modelling and visualisation (latent dirichlet allocation)
- Automatic classification using predictive modelling based on text data
- Visualisation of correlations & topics
- Word embeddings
- Document similarities & Text alignment
Hope to see you there.
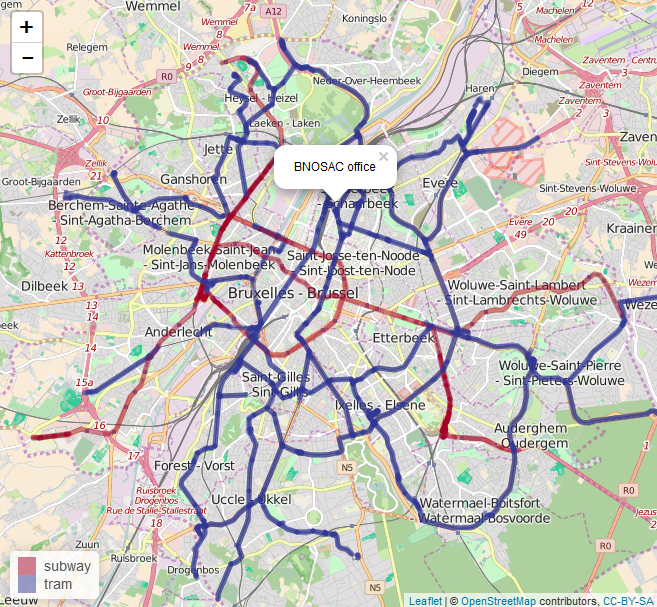
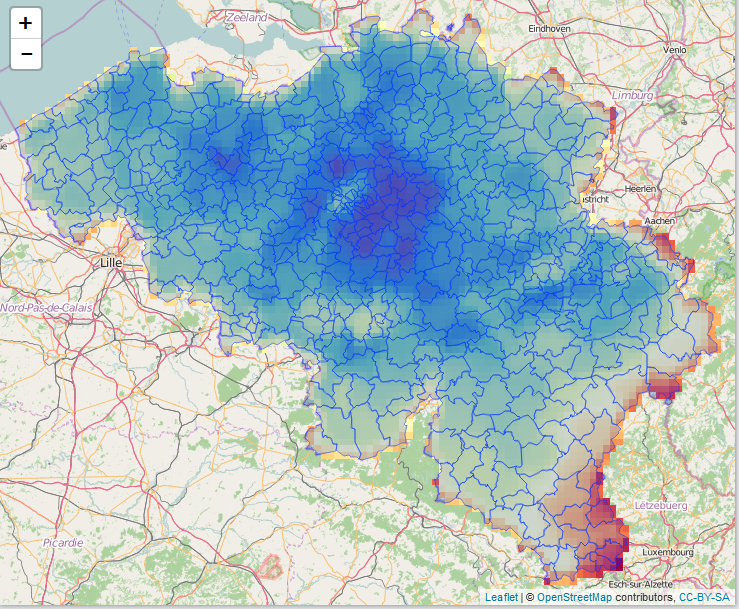
Within 2 weeks, our 2-day crash course on Applied spatial modelling with R (April 13-14, 2016) will be given at the University of Leuven, Belgium: https://lstat.kuleuven.be/training/applied-spatial-modelling-with-r
You'll learn during this course the following elements:
- The sp package to handle spatial data (spatial points, lines, polygons, spatial data frames)
- Importing spatial data and setting the spatial projection
- Plotting spatial data on static and interactive maps
- Adding graphical components to spatial maps
- Manipulation of geospatial data, geocoding, distances, …
- Density estimation, kriging and spatial point pattern analysis
- Spatial regression
More information: https://lstat.kuleuven.be/training/applied-spatial-modelling-with-r. Registration can be done at https://lstat.kuleuven.be/forms/courses
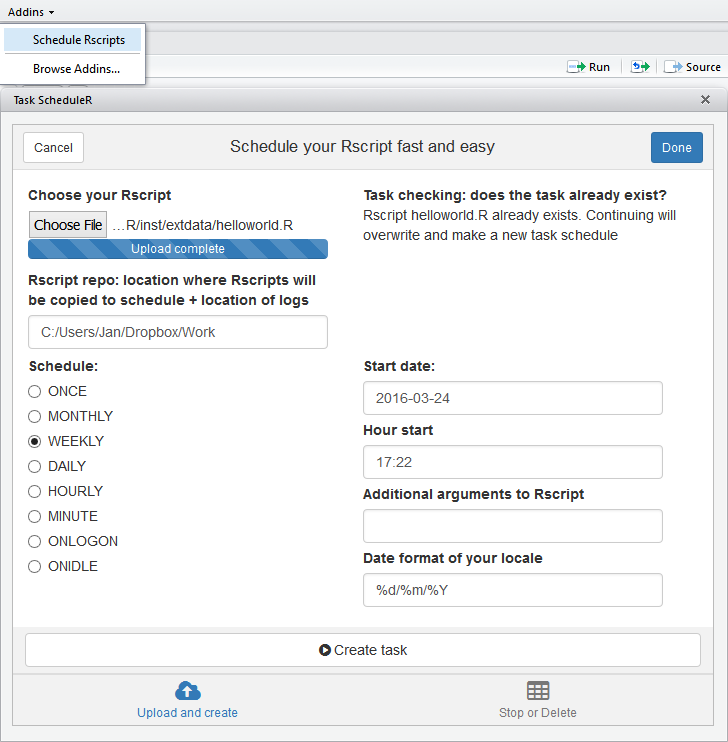
With the release of RStudio add-in possibilities, a new area of productivity increase and expected new features for R users has arrived. Thanks to the help of Oliver who has written an RStudio add-in on top of taskscheduleR, scheduling and automating an R script from RStudio is now exactly one click away if you are working on Windows.
How? Just install these R packages and you have the add-in ready at the add-in tab in your RStudio session. Select your R script and schedule it to run any time you want. Hope this saves you some day-to-day time and feel free to help make additional improvements. More information: https://github.com/bnosac/taskscheduleR.
install.packages('data.table')
install.packages('knitr')
install.packages('miniUI')
install.packages('shiny')
install.packages("taskscheduleR", repos = "http://www.datatailor.be/rcube", type = "source")
If you are working on a Windows computer and want to schedule your R scripts while you are off running, sleeping or having a coffee break, the taskscheduleR package might be what you are looking for.

The taskscheduleR R package is available at https://github.com/bnosac/taskscheduleR and it allows R users to do the following:
i) Get the list of scheduled tasks
ii) Remove a task
iii) Add a task
- A task is basically a script with R code which is run through Rscript
- You can schedule tasks 'ONCE', 'MONTHLY', 'WEEKLY', 'DAILY', 'HOURLY', 'MINUTE', 'ONLOGON', 'ONIDLE'
- After the script has run, you can check the log which can be found at the same folder as the R script. It contains the stdout & stderr of the Rscript.
Below, you can find an example how you can schedule your R script once or daily in the morning.
library(taskscheduleR)
myscript <- system.file("extdata", "helloworld.R", package = "taskscheduleR")
## run script once within 62 seconds
taskscheduler_create(taskname = "myfancyscript", rscript = myscript,
schedule = "ONCE", starttime = format(Sys.time() + 62, "%H:%M"))
## run script every day at 09:10
taskscheduler_create(taskname = "myfancyscriptdaily", rscript = myscript,
schedule = "DAILY", starttime = "09:10")
## delete the tasks
taskscheduler_delete(taskname = "myfancyscript")
taskscheduler_delete(taskname = "myfancyscriptdaily")
- When the task has run, you can look at the log which contains everything from stdout and stderr. The log file is located at the directory where the R script is located.
## log file is at the place where the helloworld.R script was located
system.file("extdata", "helloworld.log", package = "taskscheduleR")
Who wants to set up an RStudio add-in for this?